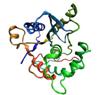
 Protein Knots
Protein Knots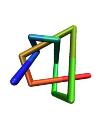
 Protein Knots
Protein Knots
List of known knots
| Protein | Species | PDB code | Length | Knot | Knotted core |
|---|---|---|---|---|---|
| YbeA-like | E.coli | 1ns5 | 153 | 31 | 69-121 (32) |
| T.maritima | 1o6d | 147 | 31 | 68-117 (30) | |
| S.aureus | 1vh0 | 157 | 31 | 73-126 (31) | |
| B.subtilis | 1to0 | 148 | 31 | 73-125 (32) | |
| tRNA(m1G37)-methyltransferase TrmD | H.influenza | 1uaj | 241 | 31 | 85-130 (92) |
| E.coli | 1p9p | 235 | 31 | 90-130 (89) | |
| S.cerevisiae | 2v3k | 219 | 31 | 175-225 (27) | |
| SpoU-like RNA 2'-O ribose mtf. | T.thermophilus | 1v2x | 191 | 31 | 96-140 (51) |
| H.influenza | 1j85 | 156 | 31 | 77-114 (42) | |
| T.thermophilus | 1ipa | 258 | 31 | 190-234 (29) | |
| E.coli | 1gz0 | 242 | 31 | 173-215 (28) | |
| A. aeolicus | 1zjr | 197 | 31 | 100-144 (58) | |
| S. viridochromog. | 1x7p | 267 | 31 | 209-251 (31) | |
| YggJ C-terminal domain-like | H.influenza | 1nxz | 246 | 31 | 166-217 (30) |
| B.subtilis | 1vhk | 235 | 31 | 168-226 (27) 1 | |
| T.thermophilus | 1v6z | 227 | 31 | 104-203 (25) 3 | |
| Hypothetical protein MTH1 (MT0001) | A.M. thermoautotr. | 1k3r | 262 | 31 | 48-234 (28) |
| Carbonic Anhydrases: | |||||
| Carbonic Anhydrase | N. gonorrhoeae | 1kop | 223 | 31 | 39-226 (0) |
| Carbonic Anhydrase I | H.sapiens | 1hcb | 258 | 31 | 29-256 (2) |
| Carbonic Anhydrase II | H.sapiens | 1lug | 259 | 31 | 31-257 (3) |
| Bos Taurus | 1v9e | 259 | 31 | 32-256 (3) | |
| Dunaliella salina | 1y7w | 274 | 31 | 39-272 (4) | |
| Carbonic Anhydrase III | Rattus norv. | 1flj | 259 | 31 | 31-258 (3) |
| H. sapiens | 1z93 | 263 | 31 | 31-258 (9) | |
| Carbonic Anhydrase IV | H.sapiens | 1znc | 262 | 31 | 31-258 (1) |
| Carbonic Anhydrase V | Mus musculus | 1keq | 238 | 31 | 31-258 (4) |
| Carbonic Anhydrase VII | H.sapiens | 1jd0 | 260 | 31 | 31-258 (3) |
| Carbonic Anhydrase XIV | Mus Musculus | 1rj6 | 259 | 31 | 31-258 (2) |
| Miscellaneous: | |||||
| Ubiquitin Hydrolase UCH-L3 | H.sapiens | 1xd3 | 229 | 52 | 13-212 (12) 4 |
| S.cerevisiae (synth.) | 1cmx | 214 | 31 | 14-228 (6) 4,1 | |
| Ubiquitin Hydrolase UCH-L1 | H.sapiens | 2etl | 219 | 52 | 10-216 (13) |
| S-adenosylmethionine synthetase | E.coli | 1fug | 383 | 31 | 33-260 (32) |
| rattus norv. | 1qm4 | 368 | 31 | 46-281 (29) 1 | |
| H. sapiens | 2p02 | 380 | 31 | 59-302 (21) | |
| Class II ketol-acid reductoisomerase | Spinacia oleracea | 1yve | 513 | 41 | 321-533 (62) |
| E.coli | 1yrl | 487 | 41 | 222-437 (52) 2 | |
| Transcarbamylase | B.fragilis | 1js1 | 324 | 31 | 169-267 (57) |
| X.campestris | 1yh0 | 328 | 31 | 173-277 (57) 2 | |
| Methyltransferase | H.sapiens | 2ha8 | 159 | 31 | 103-148 (30) |
| P. gingivalis | 2i6d | 231 | 31 | 177-223 (9) |
To speed up calculations, the KMT reduction scheme is used. This algorithm successively deletes amino acids that are not essential to the topological structure of the protein. It is also employed to create a reduced representation of the knot. In the course of our investigations we came up with a number of stringent criteria that a structure should satisfy to be classified as knotted:
Unfortunately, there are some structures containing regions of the backbone that were not resolved and for which coordinates are not reported in PDB (a gap in the structure). Mobile loops may not be resolved by X-ray crystallography unless they are stabilized by a ligand or by protein engineering, for example. If the polypeptide chain contains a gap, the knot is reported if a knot is present in at least one fragment of the chain and the structure that results from gaps being bridged with straight lines contains a knot. These criteria form the basis of our list of known knots. We have also included knotted structures with gaps if at least one homolog is knotted.
Questions about particular knots? Contact Peter Virnau.